|
If the coverage is less than what you specified, CodonCode Aligner can change the consensus base to 'N', 'X', or '-'. For example, a 100,000 bases long genomic reference sequence can be clipped to single exon regions after an alignment.Īutomatically mask consensus bases in regions with low coverage. This new feature allows automatic clipping of very long reference sequences to the region you want to look at. Choose from clipping the reference sequence to a number of uncovered bases on each side, clipping to the alignment or the exon, or not clipping at all. Uncovered reference sequences can now be shortened automaticaly after an alignment in CodonCode Aligner. Easily identify regions with many differences, then zoom for a closer look.Ĭhanging the zoom level directly through the popup menu at the bottom left corner of the aligned bases is fast and easy.Īutomatically shorten reference sequence after alignments Use highlighting of discrepancies and ambiguities to easily spot preserved regions. Zoom out to view many sequences and their differences at once. You can export the phylogenetic trees in Newick format. View your trees with branch lengths that represent the number of changes or see the topolgy only. Each tree is displayed right next to your samples in the contig view. Rebuilding the tree after edits in a contig can help you to verify the changes. 
You can use the phylogenetic trees as quality control for a simple way to find unexpected differences between your sequences and expected results. Automatically sort your samples by distance when building a tree. Version 3.7.1 added the option to replace '-' characters in consensus sequences with 'n', and included a number of important bug fixes.īuild Neighbor-Joining trees for your contigs in CodonCode Aligner. Note: The START button will be disabled if the Job list contains less than two files.CodonCode Aligner version 3.5 introduced several new features and improvements like building Neighbor-Joining trees, zooming in the contig view, shortening reference sequences after alignments, masking contig bases in low coverage regions, the percentage consensus as additional consensus method, and much more. To delete files from the Job List you can use Delete selected and DeleteĪlternatively, you can use the keyboard shortcut by pressing the Delete key. 
To add the files in the Job List you can either drag and drop or use the buttons Add file and Add Types to be displayed (for a description of the file types supported by DNA Baser, please visit this page). From the File Type filter box, you can choose what file The files will appear in the File List box. 
In the Job List box are the files that are going to be assembled.įrom the Folder List panel select the folder were your sequence files are. Soft: bioedit, FinchTVĪb ab1 abi scf seq fasta, academic, align, article, aligner, alignment, assemble, assembler, assembling, base, cheap, code, computer, consensus, contig, conversion, convert, convertor, detection, dna assembly software, free download, freeware, gap, mac os x, linux, windows xp, mutation, price, primer, real-time pcr, quality, research, sequence, Sequencing, trace, traces, sequencher codoncode vector nti alternative, ztrĮxploring for files. 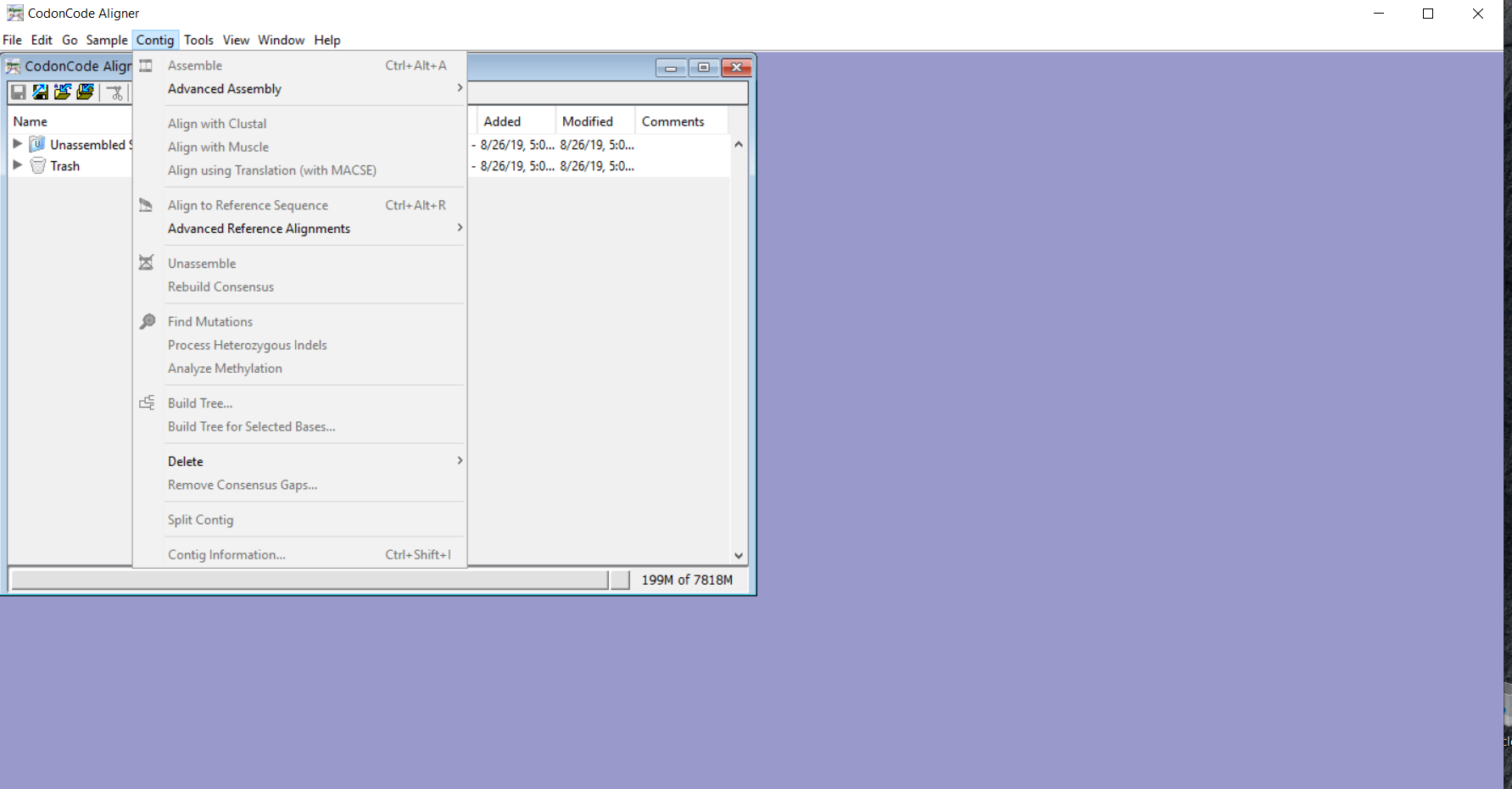
Similar expensive software: Sequencher, SeqMan 2, ChromasPro CodonCode, Invitrogen Vector NTI, phrap phred. An affordable alternative to the really expensive software for DNA assembly and alignment.ĭNA BASER can import/open scf, gel, ab1, abi, fasta, seq, sequence file for dynamic alignment.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
